Background:
The base composition of Whole Genome Bisulfite Sequencing (WGBS) libraries exhibits a severe imbalance (GC content of approximately 20%), posing significant challenges for delivering high-quality WGBS sequencing data. Conventionally, this issue is mitigated by incorporating high proportions of specialized balanced libraries (e.g., PhiX or E. coli libraries) or co-sequencing with samples featuring 40%-50% GC content. However, these methods reduce production efficiency and increase sequencing costs and operational complexity.
Methylation sequencing plays a pivotal role in early tumor screening by detecting epigenetic alterations that precede genetic mutations. It offers a non-invasive and highly sensitive approach for cancer detection at its earliest stages. GeneMind high-throughput sequencing platforms(GenoLab M, FASTASeq 300, and SURFSeq 5000) have achieved high-quality sequencing with “0% Phix library spike in.” In this test, under 0% PhiX spike in conditions, the system demonstrated a mean Q30 ≥92% and a mean single-index demultiplexing rate ≥98%, delivering data performance comparable to the competitor NV platform when mixed with balanced libraries.
Experiment Design:
| Sequencing Platform | Proportion of PhiX Spike in | Repeats | Sample ID |
| FASTASeq 300 | 0% | 3 | 1_FA, 2_FA, 3_FA |
| 20% PhiX | 1 | PhiX_FA | |
| GenoLab M | 0% | 3 | 1_GL, 2_GL, 3_GL |
| 20% PhiX | 1 | PhiX_GL | |
| SURFSeq 5000 | 0% | 3 | 1_SF, 2_SF, 3_SF |
| Competitor NV | 80% Phix | 3 | 1_NV, 2_NV, 3_NV |
Result:
(1)Achieved a mean Q30 ≥92% and a mean single-index demultiplexing rate ≥98%;
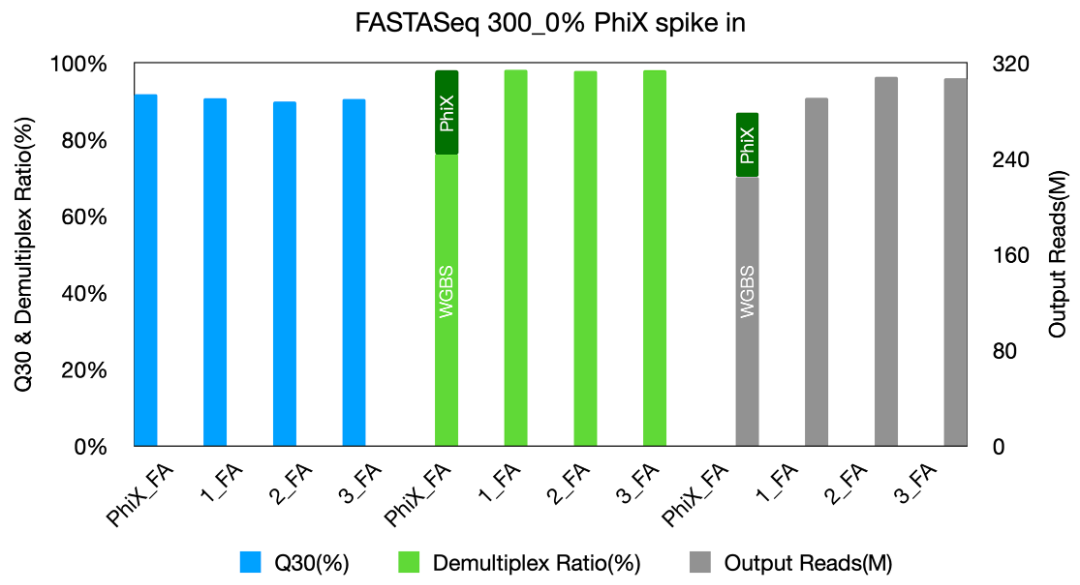
Fig 1: Data performance of FASTASeq 300_0% PhiX spike in
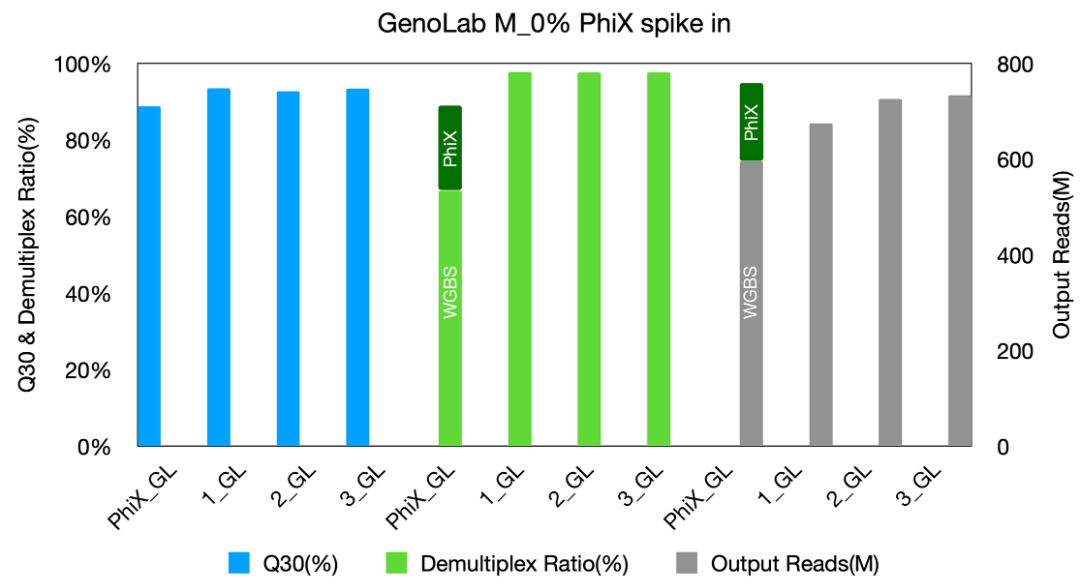
Fig 2: Data performance of GenoLab M_0% PhiX spike in
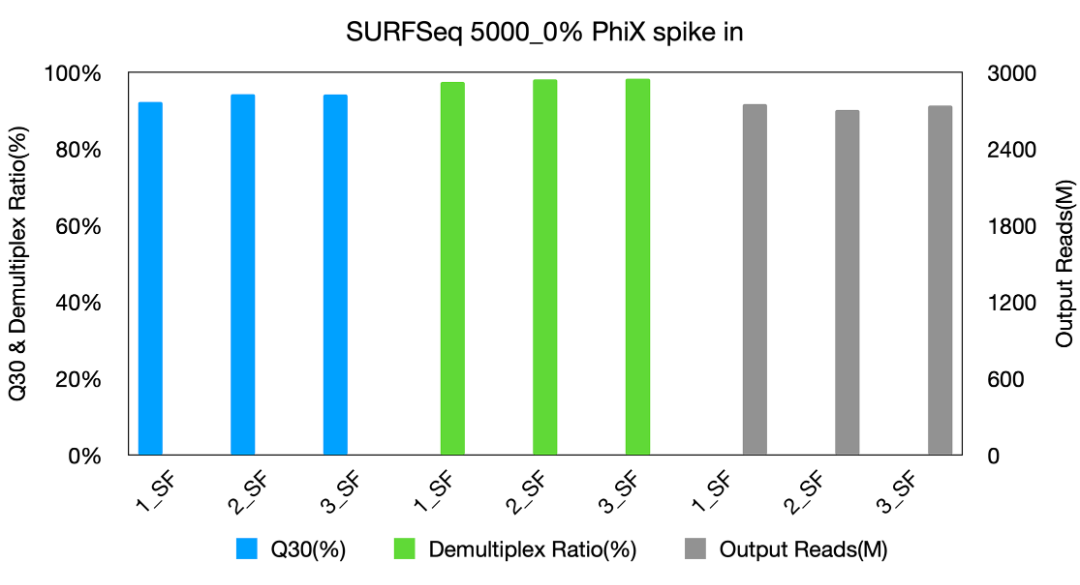
Fig 3: Data performance of SURFSeq 5000_0% PhiX spike in
(2) Demonstrated excellent cross-platform data performance, exceeding 80%;
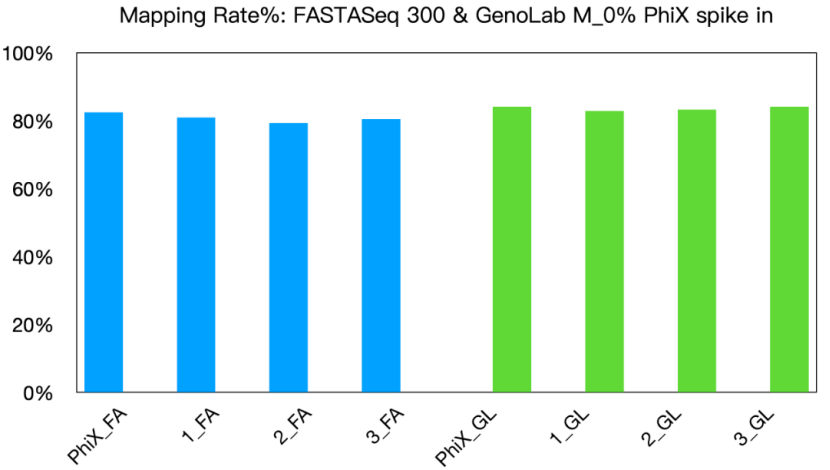
Fig 4: Mapping rate in FASTASeq 300 and GenoLab M platform
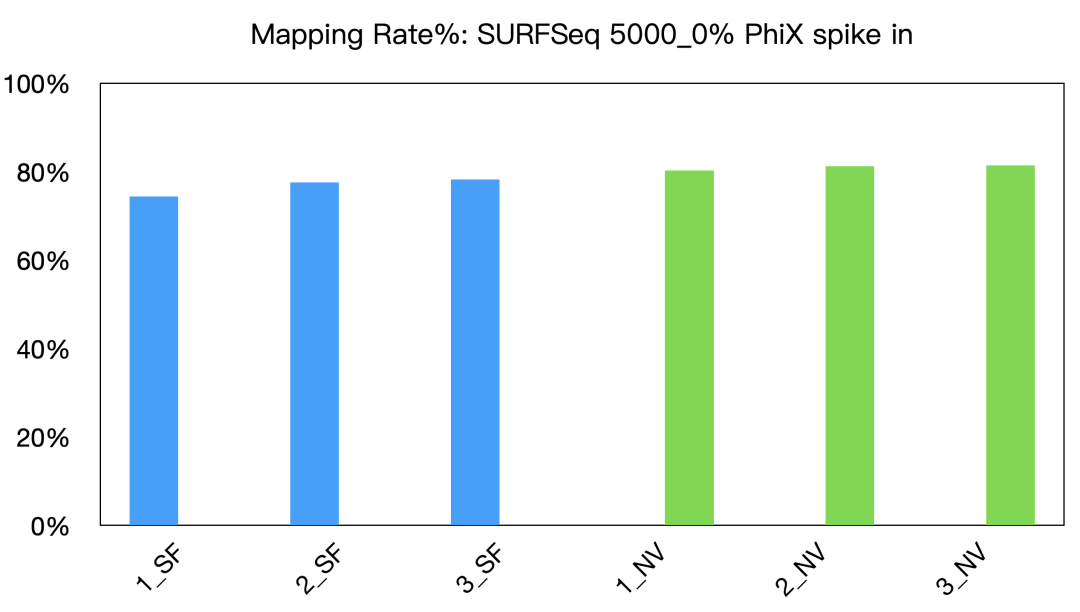
Fig 5: Mapping rate in SURFSeq 5000 and competitor NV platform
(3)Maintained highly consistent distribution ratios of methylation types and methylation level profiles in the whole genome.
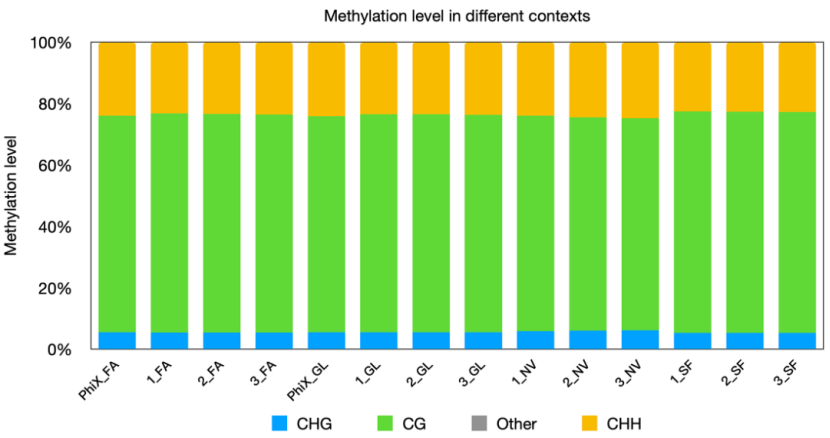
Fig 6. Distribution ratios of methylation types
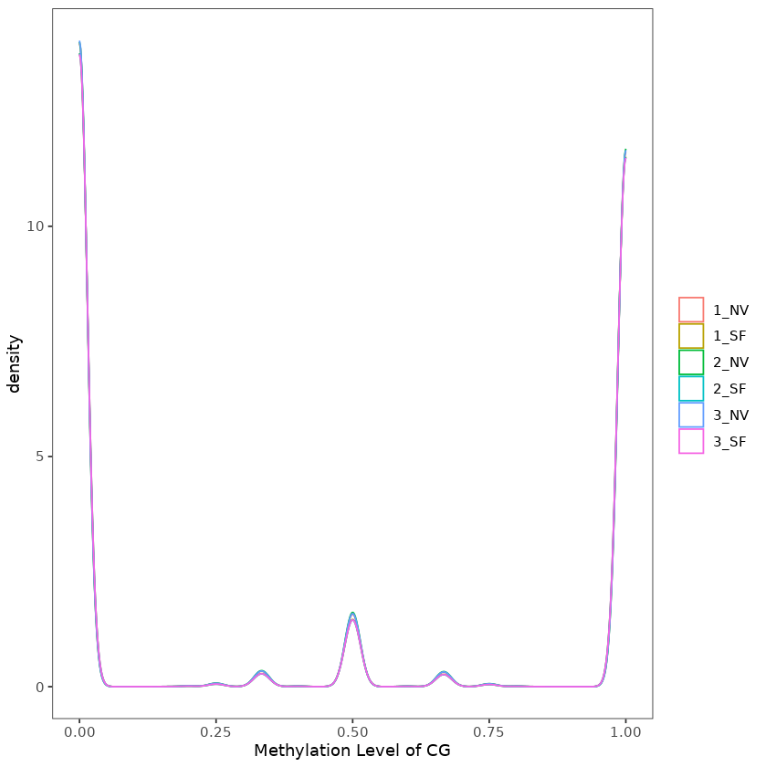
Fig 7: Methylation level of CG in whole genome
Conclusion:
GeneMind has achieved high-quality WGBS sequencing data with “0% PhiX library spike in” across all NGS product lines through technology optimization and upgrades. This breakthrough has further enhanced the product competitiveness of GeneMind sequencing platforms by providing end-users with highly efficient, cost-effective sequencing solutions.




